Why did we develop SIC?
In cheminformatics,molecular fingerprints have long been used to evaluate structural similarity.They are powerful and widely adopted,but they also have an important limitation:they often struggle to represent very small structural differences in a continuous and interpretable way,especially within groups of structural isomers.When compounds share the same molecular formula,their differences can be subtle,yet biologically and chemically meaningful.Conventional methods do not always capture these differences clearly.
This was the motivation behind the development of Structural Isomer Cumulative molecular fingerprint,or SIC.Instead of relying only on overall molecular structure patterns,SIC focuses on substructural differences and the molecular weights associated with those differences.By accumulating these differences within compounds that share the same molecular formula,SIC enables a relative and explicit evaluation of structural distance.
What problem does SIC solve?
Traditional similarity metrics are often affected by factors that are not directly related to the structural question we want to ask.Two of the biggest issues are dataset size and molecular complexity.In practice,this means that similarity distributions can shift depending on how many compounds are included in the analysis or how large and complex the molecules are.As a result,it becomes difficult to compare structural distances consistently across datasets.
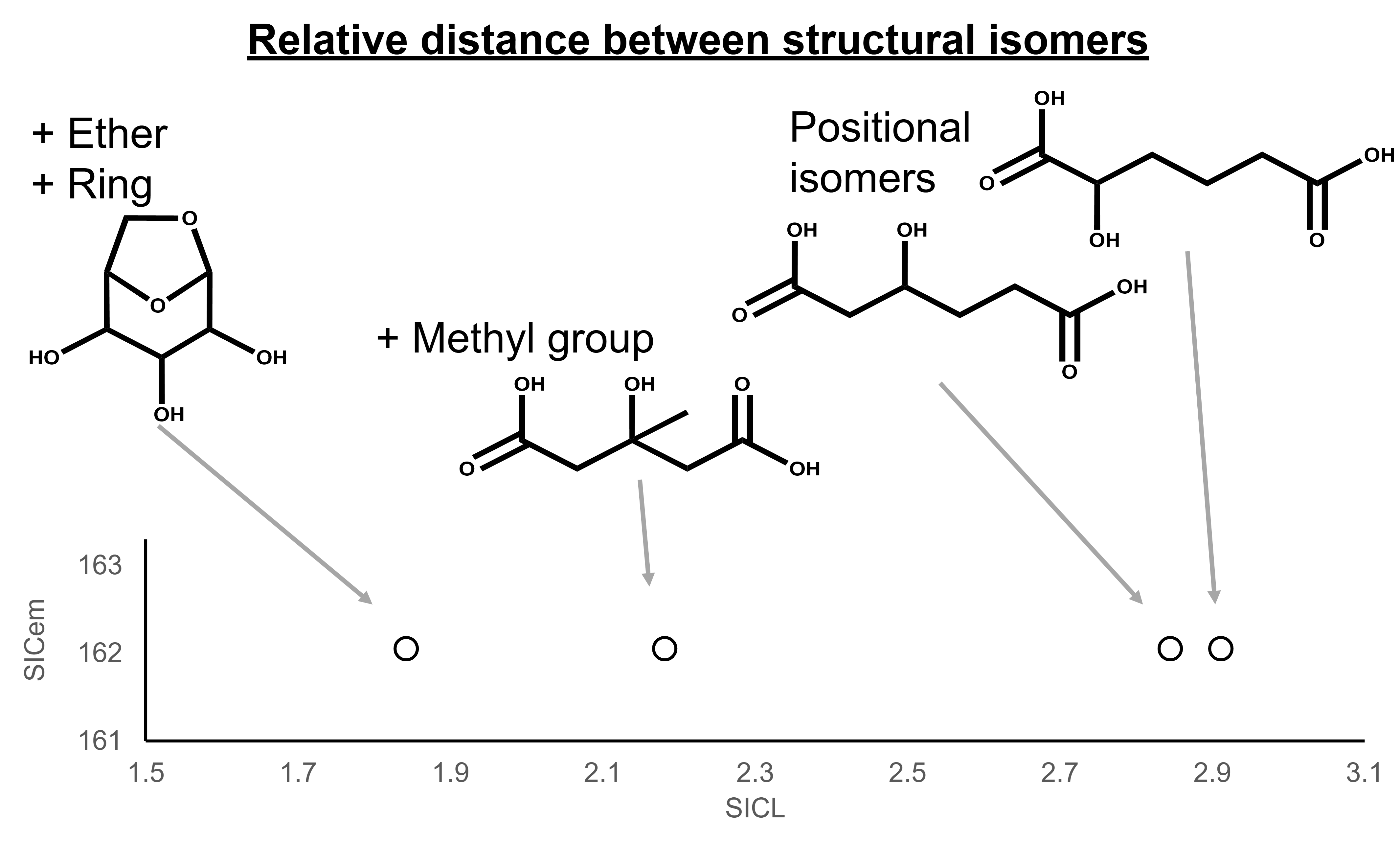
SIC was designed to reduce this problem.Its structure-based variables,including SICL and SICem,make it possible to visualize chemical space in a way that is easier to interpret.In our analyses,SICem reflected molecular weight variation,while SICL represented substructural differences.This separation allowed the compound distribution to be organized more clearly than with conventional fingerprints.Even positional isomers,which are notoriously difficult to distinguish with standard methods,could be relatively evaluated as distinct points in structural space.
How was SIC evaluated?
To test whether SIC actually improves structural evaluation,we compared it against conventional molecular fingerprints using several structural isomer datasets obtained from PubChem.These included molecular formula groups with very different dataset sizes and molecular weights,such as C₆H₆O₂ with 377 compounds and C₄₈H₈₉NO₁₈ with 31 compounds.We also benchmarked SIC across five isomer groups in total and visualized the resulting chemical spaces using principal component analysis.
The purpose of this comparison was not simply to show that SIC behaves differently.Rather,it was to determine whether SIC provides a more stable and interpretable representation of structural difference when conventional methods become sensitive to dataset scale or molecular weight.The results showed that SIC reduced the influence of both factors while preserving fine-grained structural resolution.
Beyond isomers:generalizability and toxicological screening
We also wanted to know whether SIC would remain useful outside a narrow benchmark setting.For this reason,we applied the method to diverse metabolite databases derived from both prokaryotic and eukaryotic organisms,including HMDB,YMDB,and ECMDB,as well as the toxic compound database T3DB.Across these datasets,SICem and SICL clearly represented structural distributions in relation to both molecular weight and structural diversity.
One particularly interesting result emerged in toxicity-related analyses.Using SIC,we were able to identify many endogenous metabolites that were structurally close to known toxic compounds,including relationships that were not captured by previous methods.This suggests that SIC may offer a useful framework for toxicological screening and for exploring chemical neighborhoods that are relevant to biological risk.
What makes SIC different?
The key contribution of SIC is that it provides an explicit structural distance rather than a score that easily becomes redundant or saturated.Conventional chemical space visualization methods often show clusters,but do not clearly explain which dimensions correspond to which structural properties.SIC addresses this by using two interpretable fingerprint variables.This makes structural diversity easier to read and helps researchers understand why compounds are positioned where they are.
Why SIC matters for BCC
At BCC,we aim to build tools that make scientific interpretation more explicit,practical,and reproducible.SIC fits this philosophy well.It does not treat structural similarity as a black box.Instead,it offers a framework for understanding how structural differences accumulate and how those differences shape chemical space.
SIC is not just another fingerprint.It is an attempt to ask a more precise question:how should we measure structural difference when tiny changes matter? For structural isomers,toxicological screening,and diversity-aware chemical analysis,that question is more important than ever.
We currently provide SIC through BCC,where users can explore the standard implementation here:https://biochemcalc.com/nor_sic.
For users who need extended functionality,we also distribute a paid version called extend SIC (ESIC) through AMEZIST.More information is available here:https://www.tils.co.jp/amezist.
By offering both the standard SIC implementation in BCC and the extended ESIC version in AMEZIST,we hope to support a broader range of researchers working on structural comparison,compound interpretation,and chemical space analysis.
Paper information
The SIC method is described in our research article titled Structural Isomer Cumulative molecular fingerprinting method (SIC) for standardizing structural isomeric relationships.This paper was accepted in Communications Chemistry on 2025-11-06.The DOI is 10.1038/s42004-025-01798-3.